
Elhanan Borenstein, Ph.D.

Associate Professor

Blavatnik School of Computer Science

and

Sackler Faculty of Medicine

Tel Aviv University
Affiliate Professor
Dept of Genome Sciences
University of Washington
External Professor
Santa Fe Institute
|
 |
 |
 |
| Download |
 |
 | Datasets from published papers:
|
 |
 |
 |
Defining and Evaluating Microbial Contributions to Metabolite Variation in Microbiome-Metabolome Association Studies
Noecker et al., mSystems, 2019.
» Simulation results and associated data (zip)
» FBA co-culture simulation code and configuration files (zip)
|
 |
 |
 |
 |
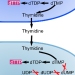
Metabolic Model-Based Analysis of the Emergence of Bacterial Cross-Feeding via Extensive Gene Loss
McNally and Borenstein, BMC Systems Biology, 2018.
» Gene content and order of deletions of evolved two-species communities (zip)
|
 |
 |
 |
 |
Evolutionary assembly patterns of prokaryotic genomes
Press et al., Genome Research, 2016.
» Final PGCE dependency network (xlsx)
» Parameter file for gainLoss run (txt)
» Log file for gainLoss run (txt)
» Code and data archive (zip)
|
 |
 |
 |
 |
Metabolic model-based integration of microbiome taxonomic and metabolomic profiles elucidates mechanistic links between ecological and metabolic variation
Noecker et al., mSystems, 2016.
» Code archive (zip) - for the latest version of this method, see our MIMOSA software page
» Dataset 1: (qPCR-based community composition, metabolomics)
» Dataset 2: (16S-based community composition, metabolomics)
» Dataset 3: (16S-based community composition, metabolomics)
» Dataset 4: (Shotgun-based functional composition, metabolomics, metabolite IDs)
|
 |
 |
 |
 |
A large-scale analysis of strain-level copy number variation in the human gut microbiome
Greenblum et al., Cell, 2015.
» Code archive for mapping and analysis pipelines (zip)
» Copy number estimates of 40,088 KCs across 109 samples (excel)
» A list of highly and set-specific variable KCs (excel)
|
 |
 |
 |
 |
Emergent biosynthetic capacity in simple microbial communities
Chiu et al., PLoS Computational Biology, 2014.
» Dataset 1 (6 manually curated models) - models (zip)
» Dataset 1 (6 manually curated models) - media (zip)
» Dataset 2 (116 SEED models) - models (zip)
» Dataset 2 (116 SEED models) - media (zip)
|
 |
 |
 |
 |
Reconstructing the genomic content of microbiome taxa through shotgun metagenomic deconvolution
Carr et al., PLoS Computational Biology, 2013.
» Dataset S1: simple synthetic metagenomic samples (xls)
» Dataset S2: 3 strain metagenomic samples with sequencing and annotation error (xls)
» Dataset S3: 20 strain metagenomic samples with sequencing and annotation error (xls)
» Code archive (zip)
|
 |
 |
 |
 |
Metabolic modeling of species interaction in the human microbiome elucidates community-level assembly rules
Levy and Borenstein, PNAS, 2013.
» Supplementary information file
» Dataset S1: Metabolic interaction and co-occurrence of MetaHIT microbiome species (excel)
» Dataset S2: Abundance, metabolic interaction, and co-occurrence of HMP species (excel)
|
 |
 |
 |
 |
Circuitry and dynamics of human transcription factor regulatory networks
Neph et al., Cell, 2012.
» Supplementary information file
» Dataset containing full computed networks for all cell types (zip)
» Table S1: Order of factors in all circos diagrams and hive plots (pdf)
» Table S2: Edges unique to a cell type typically form a well-connected subnetwork (pdf)
» Table S3: Sizes and statistics of derived regulatory networks (pdf)
|
 |
 |
 |
 |
Metagenomic systems biology of the human gut microbiome reveals topological shifts associated with obesity and inflammatory bowel disease
Greenblum et al., PNAS, 2012.
» Supplementary information file
» Gut microbiome enzymes with disease association and topological data (excel)
» Gut microbiome, community-level network, all samples (txt)
» Gut microbiome, community-level network, lean healthy samples (txt)
» Gut microbiome, community-level network, obese samples (txt)
» Gut microbiome, community-level network, IBD samples (txt)
» Gut microbiome, community-level network, with topological measures (Cytoscape session)
|
 |
 |
 |
 |
Large-scale reconstruction and phylogenetic analysis of metabolic environments
Borenstein et al., PNAS, 2008
» Supplementary information file
» Occurring compounds and seed set across 487 species (dataset S1, txt)
» Phylogenetic patterns and statistics for 2163 compounds (dataset S2, excel)
|
 |
 |
 |
 |
The evolution of modularity in bacterial metabolic networks, Kreimer et al., PNAS, 2008
» Supplementary information file
» Modularity and environmental information for 325 microbial species (dataset 1, excel)
|
 |

|
 |