| Software |
 |
 |
 |
 |
NetSeed
NetSeed is a software package for identifying the seed sets of networks (as defined in Borenstein et al., 2008) and is available as an online tool and as a Perl module.
|
 |
 |
 |
 |
NetCooperate
NetCooperate is a software package for determining host-microbe and microbe-microbe cooperative potential and is available as an online tool and as a Python module.
|
 |
 |
 |
 |
MetaDecon
MetaDecon is a computational framework, integrating variation in gene abundances across multiple samples with taxa abundance data to infer taxon-specific gene profiles and to reconstruct the genomic content of community members.
|
 |
 |
 |
 |
MUSiCC
MUSiCC is a software package for normalizing and correcting gene abundance measurements derived from metagenomic shotgun sequencing.
|
 |
 |
 |
 |
FishTaco
FishTaco is a computational framework for linking taxonomic and functional dynamics observed in metagenomic samples and for identifying taxa that drive disease-associated functional shifts in the microbiome.
|
 |
 |
 |
 |
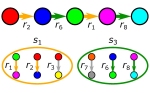
CoMiDA
CoMiDA is an algorithm for designing simple communities with some predefined metabolic capacities.
|
 |
 |
 |
 |
MIMOSA
MIMOSA is a novel framework for mechanistically linking microibome ecology and metabolomic data.
|
 |
 |
 |
 |
MIMOSA2
MIMOSA2 is an extension of MIMOSA - a novel framework for mechanistically linking microibome ecology and metabolomic data. Among other features, it provides a web-application interface, new visualization, and a new algorithm for calculating taxonomic contributions to metabolite variation.
|
 |
 |
 |
 |
EMPANADA
EMPANADA is a tool for evidence-based, non-uniform, and sample-specific assignment of gene families to pathways in metagenomic data.
|
 |
 |
 |
 |
BURRITO
BURRITO is a web-based tool for interactive exploration of metagenomic datasets, linking taxonomic and functional microbiome profiles.
|
 |
 |
 |
 |
MetaLAFFA
MetaLAFFA is a flexible, end-to-end, and compute cluster-compatible metagenomic functional annotation pipeline.
|
 |
